Use this to save results from a Finlay & Wilkinson joint regression analysis in Genstat data structures.
- After selecting the appropriate boxes, type the names for the identifiers of the data structures into the corresponding In: fields.
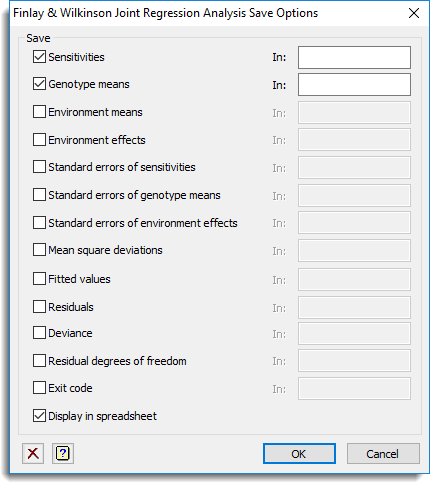
Save
| Sensitivities | Table | The estimates of sensitivities for each genotype |
| Genotype means | Table | The estimates of genotype means |
| Environment means | Table | The estimates of environment means |
| Environment effects | Table | The estimates of environment effects |
| Standard errors of sensitivities | Table | The standard errors of the sensitivities |
| Standard errors of genotype means | Table | The standard errors of genotype means |
| Standard errors of environment effects | Table | The standard errors of environment effects |
| Mean square deviations | Table | The mean square deviations about the line fitted to each genotype |
| Fitted values | Variate | The fitted values from the model |
| Residuals | Variate | The residuals from the model |
| Deviance | Scalar | The residual degrees of freedom from the fitted model |
| Residual degrees of freedom | Scalar | The residual degrees of freedom from the model |
| Exit code | Scalar | The exit status: set to 0 if the analysis converged, 1 otherwise |
Display in spreadsheet
The saved results will be displayed within new spreadsheet windows, grouped by tables (one for genotypes and one for environments), variates and scalars.
See also
- Finlay & Wilkinson joint regression analysis menu
- Options for choosing which results to display and fitting options
- AMMI menu
- RFINLAYWILKINSON procedure
- AMMI procedure in command mode