The preliminary single environment analysis runs a REML analysis twice. In the first analysis the genotypes are used as a random term and in the second analysis they are fitted as a fixed term. This dialog lets you select options for information to be displayed and the method used for each analysis.
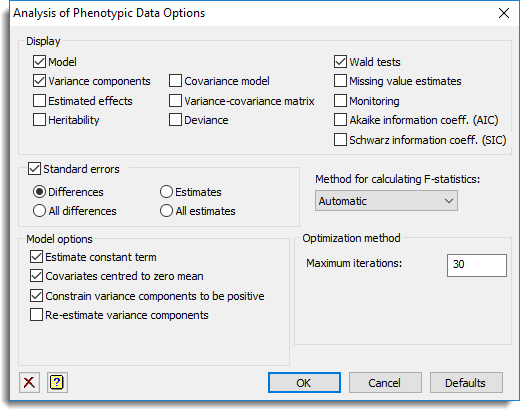
Display
This specifies which items of output are to be produced by the analysis.
| Model | Description of the model fitted by the analysis |
| Variance components | Estimates of variance parameters |
| Estimated effects | Estimates of regression coefficients for genotypes |
| Heritability | Heritability for analysis with genotypes fitted as random |
| Stratum variances | Estimates of approximate stratum variances |
| Variance-covariance matrix | Variance-covariance matrix for the variance parameters |
| Deviance | The residual deviance |
| Wald tests | Wald Tests for fixed model terms and accompanying F-statistics (if selected) |
| Missing value estimates | Estimates of values missing from the input |
| Monitoring | Monitoring information at each iteration |
| Covariance model | Estimated covariance models in matrix format |
| Akaike information coefficient (AIC) | Akaike information coefficient to assess the random model |
| Schwarz information coefficient (SIC) | Schwarz information coefficient to assess the random model |
Standard errors
Tables of effects are accompanied by estimates of standard errors. You can choose whether Genstat computes standard errors or standard errors of differences (SEDs) for the tables.
Method for calculating F-statistics
This controls whether approximate F statistics and corresponding numbers of residual degrees of freedom, are calculated in addition to Wald statistics. The computations, using the method devised by Kenward & Roger (1997), can be time consuming with large or complicated models. The default setting, automatic, can be used to allow Genstat to assess the model and decide automatically whether to do the computations and which method to use. The other settings allow you to control the computations:
| none | No F statistics are produced |
| algebraic | F statistics are calculated using algebraic derivatives (which may involve large matrix calculations) |
| numerical | F statistics are calculated using numerical derivatives (which require an extra evaluation of the mixed model equations for every variance parameter). |
Estimate missing data values
This specifies whether predictions are formed from the fitted model for missing values of the y-variate; alternatively any units with missing values in the y-variate are excluded from the analysis.
Include units with missing factor values
This specifies whether data units with missing values in any of the factors in the fixed or random models are included in the analysis. Units with missing y values are always excluded from the analysis.
Estimate constant term
Specifies whether a constant term is included in the fixed model.
Covariates centred to zero mean
Specifies whether covariates are centred to zero mean during the analysis. This applies to all covariates in the model. If covariates are centred, tables of predicted means are based on the mean covariate value, otherwise zero for each covariate.
Re-estimate variance parameters
When selected the variance parameters will be re-estimated in the second analysis. When this is not selected the variance parameters will be fixed in the second analysis using the values estimated from the first analysis when fitting genotypes in the random model.
Optimization method
The AI (Average Information) method is the standard optimization method for REML in Genstat. It uses sparse matrices and is particularly recommended for large datasets and/or complex models. An alternative method is Fisher Scoring which, if selected, also allows an absorbing factor to be specified in the model. This can be used to reduce the time or space requirements when fitting large models with many parameters. A more detailed discussion of the use and choice of absorbing factors can be found in the Genstat Reference Manual, which is available from the menu by selecting Help | Reference Manual | Directives..
Action buttons
| OK | Store the settings and close the dialog. |
| Cancel | Close the dialog without making any changes. |
| Defaults | Close the dialog without making any changes. |
Action Icons
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- REML directive for mixed model analysis in command mode